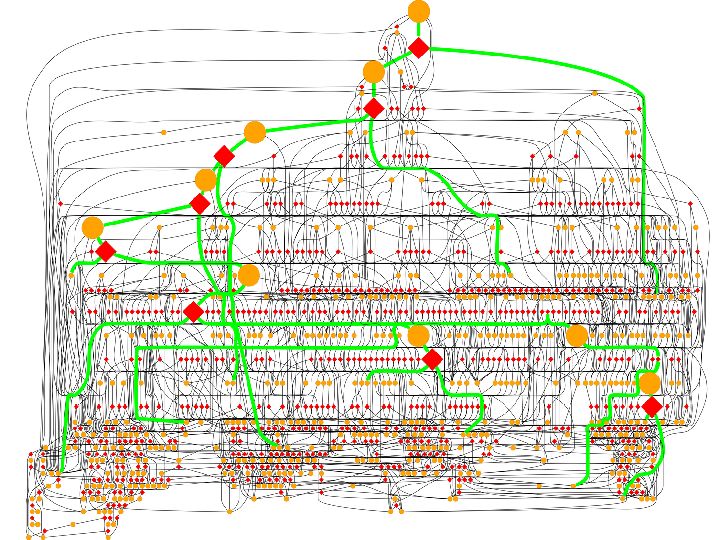
A mechanistic computational model for simulating protein unfolding.
To model the final stages of folding, and the first steps of unfolding, a mechanistic model was developed for breaking the final structure into smaller pieces via simple rotations and tranlations. The mechanistic model (GeoFold), developed in collaboration with the M. Zaki lab (Comp Sci) generates a map of all topologically possible unfolding (and implicitly, folding) pathways. Kinetic simulations can be done on this ensemble, and such simulations have been shown to reproduce observed unfolding kinetics and the effects of point mutations. Unfolding pathways generated by GeoFold are being used to understand the extreme kinetic sability of some protein, especially those studied by the W. Colon lab (RPI Chemistry).
Try the GeoFold server.